| E. coli Your Next Major Platform Programmed In Cello |
| Written by Mike James |
| Sunday, 03 April 2016 |
|
If you have grown tired of programming for desktops, phones and even IoT devices then why not move to the next level. Cello is a wetware programming language that literally lets you program bacteria.
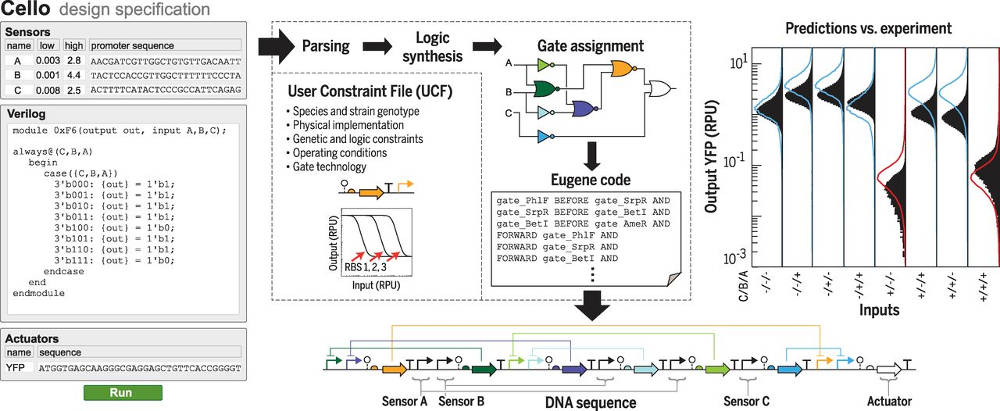
Image: Janet Iwasa Cello is based on Verilog, a hardware description language that is normally used to design digital circuits, but MIT biologists have found a way to make it create DNA encoded circuits. You write a Cello program, compile it and the target is a DNA sequence. Realize the DNA sequence put it into cell and you have the functionality you designed. The key idea is that biologists have already designed genetic modules that do specific jobs and can be assembled into working systems. The problem is that using these modules required a lot of knowledge of how they work. You could say that this is like the old days of electronic design where circuits were hand-crafted by experts who knew the characteristics of the devices that they were working with. The research team spent a lot of time designing 14 different DNA implementations of NOR/NOT logic gates that could work together in any combination without interfering with each others action. Part of the process of compiling the Verilog code is to find NOR/NOT combinations that naturally connect together to form the desired circuit. This combinatorial problem is solved using either a breadth first search or a Monte Carlo simulated annealing search. The best circuit found is then converted into one of many different DNA sequences that will produce the circuit. What is amazing is that all this seems to work. To quote from the paper reporting the results: "Cello was applied to the design of 60 circuits for Escherichia coli, where the circuit function was specified using Verilog code and transformed to a DNA sequence. The DNA sequences were built as specified with no additional tuning, requiring 880,000 base pairs of DNA assembly. Of these, 45 circuits performed correctly in every output state (up to 10 regulators and 55 parts). Across all circuits, 92% of the 412 output states functioned as predicted." The circuits were designed to measure things like the concentration of oxygen or glucose and respond. One of the circuits uses seven logic gates and at 12,000 base pairs is that largest biological circuit yet built. The diagram below shows an example circuit:
A user specifies the desired circuit function in Verilog code, and this is transformed into a DNA sequence. An example circuit is shown (0xF6); red and blue curves are predicted output states for populations of cells, and solid black distributions are experimental flow cytometry data. The outputs are shown for all combinations of sensor states; plus and minus signs indicate the presence or absence of input signal. RBS, ribosome binding site; RPU, relative promoter unit; YFP, yellow fluorescent protein. There are clearly going to be practical applications of being able to build genetic circuits so easily. Suggestions include improving yeast cells so they don't poison themselves with toxic products during fermentation, programming cells to create drugs, and so on.
Currently the software is tuned to E. coli's genetics, but it can work with other bacteria or even yeast cells. The idea is that you can write a program and then compile it to "run" on different organisms. The software is available on Github and there is a web site where you can try it out. More Informationhttps://github.com/CIDARLAB/cello Genetic circuit design automation (paywalled)
Alec A. K. Nielsen, Bryan S. Der, Jonghyeon Shin,Prashant Vaidyanathan, Vanya Paralanov, Elizabeth A. Strychalski, David Ross, Douglas Densmore, Christopher A. Voigt. Related ArticlesTraveling Salesman Applied To DNA Synthesis Sliding Blocks Are Turing Complete Edit Distance Algorithm Is Optimal Genetic Algorithms Reverse Resistance To Antibiotics Book Stored On DNA - All Knowledge In Just 4gm of DNA A New DNA Sequence Search - Compressive Genomics
To be informed about new articles on I Programmer, sign up for our weekly newsletter, subscribe to the RSS feed and follow us on, Twitter, Facebook, Google+ or Linkedin.
Comments
or email your comment to: comments@i-programmer.info |
| Last Updated ( Monday, 04 April 2016 ) |